News
Jiabei Zhu wins Best Student Paper Award in Optica Imaging Congress
Congratulations to Jiabei Zhu for winning the Best Student Paper Award in Optica Imaging Congress 3D Conference, for his paper on “3D Phase Imaging from Intensity Measurements with Non-Paraxial Multiple Scattering Model”.

Hao Wang wins Best Student Paper Award in Optica Imaging Congress
Congratulations to Hao Wang for winning the Best Student Paper Award in Optica Imaging Congress Digital Holography Conference, for his paper on "wide-field, high-resolution reflection-mode Fourier ptychographic microscopy".

CISL at Optica Imaging Congress 2023

Hao Wang
Wide-field, high-resolution reflection-mode Fourier ptychographic microscopy
* best student paper award 1st place

Jiabei Zhu
3D Phase Imaging from Intensity Measurements with Non-Paraxial Multiple Scattering Model
* best student paper award 1st place
* featured in Optica news.

Qianwan Yang
Advancing Computational Miniature Mesoscope With Simulator-Based Deep Learning Reconstruction
* featured in Optica news.

Guorong Hu
Caustic Illumination-based HiLo Microscopy

Lei Tian
Computational 3D Phase Imaging by Intensity Diffraction Tomography * invited

Lei Tian
Computational Miniature Mesoscope: augmenting miniature optics with algorithms for large-scale 3D fluorescence imaging * invited
Excited to attend CZI Imaging 2023 Annual Meeting
Full CZI Imaging program group

Scialog Fellows

Shiyi defended PhD!

Congratulations, Dr. Cheng!
=============================
Title: Augmenting Label-free Imaging Modalities with Deep Learning based Digital Staining
Presenter: Shiyi Cheng
Date: Monday, July 10th, 2023
Time: 11:00 am to 1:00 pm
Location: 8 Saint Mary's Street, Room 339
Advisor: Professor Lei Tian
Chair: Professor Ari Trachtenberg
Committee: Professor Lei Tian, Professor Eshed Ohn-Bar, Professor David A. Boas, Professor Irving Bigio, Professor Ji-Xin Cheng.
Abstract:
Label-free imaging techniques provide valuable insights into biological samples and processes in their native states, eliminating the need for labor-intensive and potentially disruptive processes of physical staining. However, these methods often lack structural and molecular specific information. To overcome this limitation, recent advances in deep learning based digital staining techniques have shown the ability to virtually introduce digital labels or stains into label-free images, which enables extracting rich information that would typically require physical staining. The integration of label-free imaging and digital staining holds great potential for significantly expanding the toolkit for biomedical imaging, facilitating improved analysis, and enhancing our understanding of biomedical sciences at both the cellular and tissue level. In this thesis, I explore supervised and semi-supervised methodologies for digital staining and their applications in augmenting label-free imaging, with a focus on imaging cytometry and human brain imaging.
In the first part of the thesis, I present a novel integration of multi-contrast dark-field reflectance microscopy and digital staining by supervised deep learning. This method enables multiplexed immunofluorescence labeling of subcellular features and single cell cytometry. By leveraging the rich structural information and sensitivity of reflectance microscopy, the digital staining method accurately predicts subcellular features and achieves up to 3 times improvement in prediction accuracy over the state-of-the-art techniques. Additionally, the method accurately reproduces single-cell level structural phenotypes related to cell cycles. The multiplexed digital labeling enables multi-parametric single-cell profiling across a large cell population.
In the second part, I developed a novel semi-supervised digital staining technique for serial sectioning OCT (S-OCT) for 3D histological imaging of human brain tissue. The deep learning model integrates unpaired image translation, a biophysical model, and unsupervised cross-modality image registration. The digital staining model enables the translation of S-OCT images to Gallyas silver staining, provides consistent staining quality across different samples, and enhances contrast across cortical layer boundaries, enabling reliable layer differentiation. Importantly, the integration of S-OCT and digital staining allows volumetric histological imaging while preserving complex 3D geometry on centimeter-scale brain tissue blocks. In addition, our pilot study demonstrates promising results on other anatomical regions acquired from different S-OCT systems.
In summary, I investigated deep-learning-based digital staining techniques for augmenting two types of label-free imaging modalities. I showcased two important applications in the field of single-cell immunofluorescence microscopy and mesoscale 3D histological human brain imaging. I expect two major potential impacts from my thesis work. First, the integration of digital staining techniques with multi-contrast microscopy can potentially enhance the throughput of single-cell imaging cytometry and phenotyping. Second, the integration of digital staining techniques with S-OCT can potentially enable high-throughput human brain imaging, facilitating comprehensive studies on the brain's structure and function. Through this exploration, this thesis advances the digital staining technique and its applications for various biomedical disciplines.
Lei is selected as a Fellow of Scialog: Advancing BioImaging


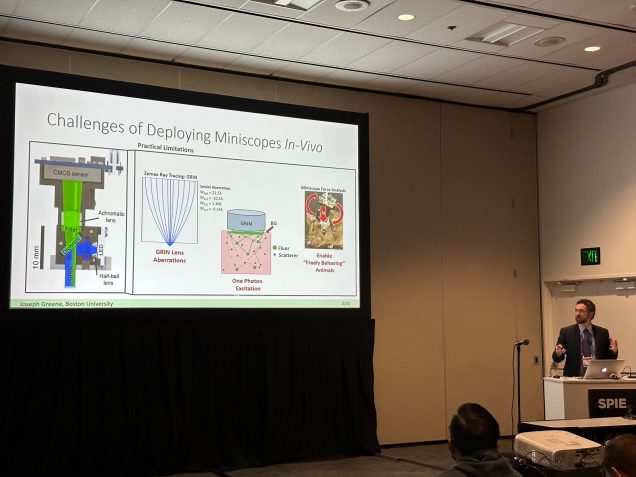
Joe, Chang present at SPIE Photonics West


Qianwan Yang wins Emil Wolf Outstanding Student Paper Award
Qianwan Yang wins Emil Wolf Outstanding Student Paper Award in Optica Frontier in Optics Congress for her work on "Computational Miniature Mesoscope with deep learning reconstruction". Congratulations!
Joe, Qianwan, Jiabei, Hao present at Optica FiO conference

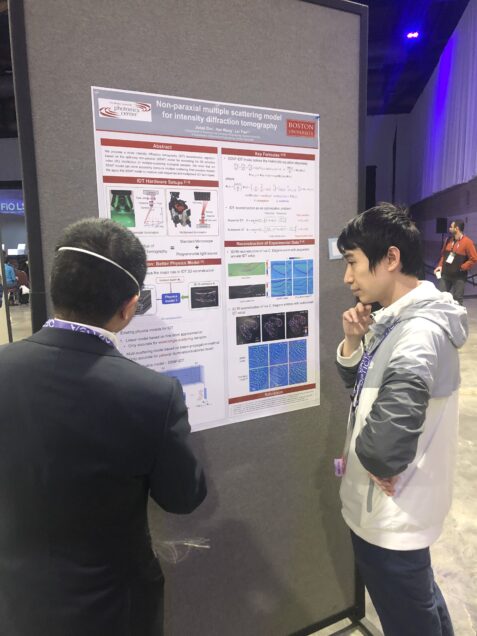
J. Zhu*, H. Wang*, L. Tian, “Non-paraxial multiple scattering model for intensity diffraction tomography”, Optica Frontier in Optics, Oct. 2022.
J. Greene*, Y. Xue*, J. Alido*, A. Matlock*, G. Hu*, K. Kilic, I. Davison, L. Tian, “Pupil engineering in miniscopes for extended depth-of-field neural imaging”, Optica Frontier in Optics, Oct. 2022.
Q. Yang*, Y. Xue*, G. Hu*, L. Tian, “Computational Miniature Mesoscope with deep learning reconstruction”, Optica Frontier in Optics, Oct. 2022. * Emil Wolf Outstanding Student Paper Award
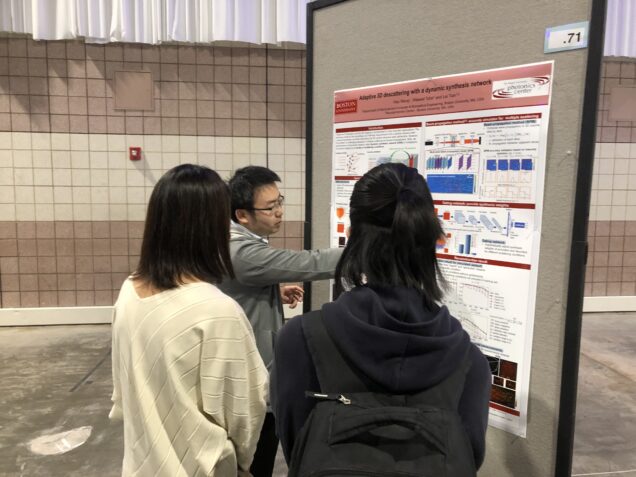
H. Wang*, W. Tahir*, L. Tian, “Adaptive volumetric descattering in digital holography”, Optica Frontier in Optics, Oct. 2022.


Yunzhe defended PhD Dissertation!

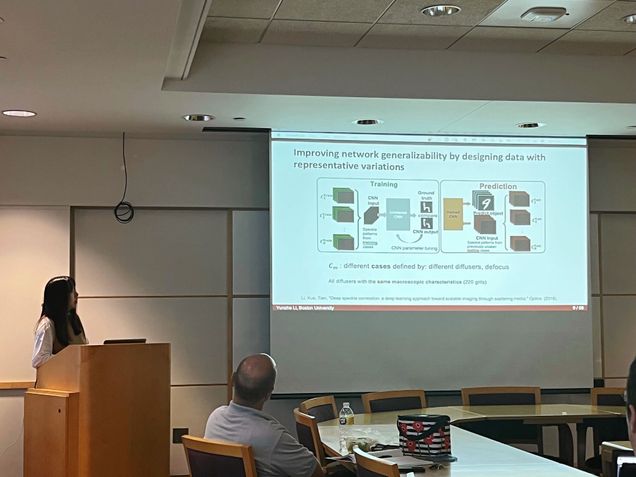
Title: Robust Deep Learning for Computational Imaging through Random Optics
Presenter: Yunzhe Li
Date: Monday, September 26, 2022
Time: 12:00 pm to 2:00 pm
Location: 8 Saint Mary's Street, Room 339
Advisor: Professor Lei Tian, ECE
Chair: Professor Eshed Ohn-Bar, ECE
Committee: Professor Lei Tian, ECE; Professor Vivek Goyal, ECE; Professor Luca Dal Negro, ECE; Professor Roberto Paiella, ECE.
Abstract: Light scattering is a pervasive phenomenon that poses outstanding challenges in both coherent and incoherent imaging systems. The output of a coherent light scattered from a complex medium exhibits a seemingly random speckle pattern that scrambles the useful information of the object. To date, there is no simple solution for inverting such complex scattering. Advancing the solution of inverse scattering problems could provide important insights into applications across many areas, such as deep tissue imaging, non-line-of-sight imaging, and imaging in degraded environment. On the other hand, in incoherent systems, the randomness of scattering medium could be exploited to build lightweight, compact, and low-cost lensless imaging systems that are applicable in miniaturized biomedical and scientific imaging. The imaging capability of such computational imaging systems, however, are largely limited by the ill-posed or ill-conditioned inverse problems, which typically causes imaging artifacts and degradation of the image resolution. Therefore, mitigating this issue by developing modern algorithms is essential for pushing the limits of such lensless computational imaging systems.
In this thesis, I focus on the problem of imaging through random optics and present two novel deep-learning (DL) based methodologies to overcome the challenges in coherent and incoherent systems: 1) no simple solution for inverse scattering problem and lack of robustness to scattering variations; and 2) ill-posed problem for diffuser-based lensless imaging.
In the first part, I demonstrate the novel use of a deep neural network (DNN) to solve the inverse scattering problem in a coherent imaging system. I propose a statistical `one-to-all' deep learning technique that encapsulates a wide range of statistical variations for the model to be resilient to speckle decorrelations. I push the limit of robustness against a broad class of perturbations including scatterer change, displacements, and system defocus up to 10X depth of field.
In the second part, I consider the utility of the random light scattering to build a diffuser-based computational lensless imaging system and present a generally applicable novel DL framework to achieve fast and noise-robust color image reconstruction. I developed a diffuser-based computational funduscope that reconstructs important clinical features of a model eye. Experimentally, I demonstrated fundus image reconstruction over a large field-of-view (FOV) and robustness to refractive error using a constant point-spread-function. Next, I present a physics simulator-trained, adaptive DL framework to achieve fast and noise-robust color imaging. The physics simulator incorporates optical system modeling, the simulation of mixed Poisson-Gaussian noise, and color filter array induced artifacts in color sensors. The learning framework includes an adaptive multi-channel L2-regularized inversion module and a channel-attention enhancement network module. Both simulation and experiments show consistently better reconstruction accuracy and robustness to various noise levels under different light conditions compared with traditional L2-regularized reconstructions.
Overall, this thesis investigated two major classes of problems in imaging through random optics. In the first part of the thesis, my work explored a novel DL-based approach for solving the inverse scattering problem and paves the way to a scalable and robust deep learning approach to imaging through scattering media. In the second part of the thesis, my work developed a broadly applicable adaptive learning-based framework for ill-conditioned image reconstruction and a physics-based simulation model for computational color imaging.