Deep learning for physics-based imaging
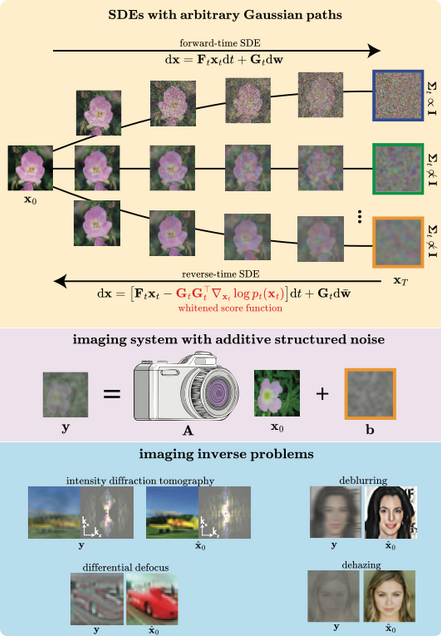
Whitened Score Diffusion: A Structured Prior for Imaging Inverse Problems
J Alido, T Li, Y Sun, L Tian
NeurIPS 2025.
⭑ Github Project
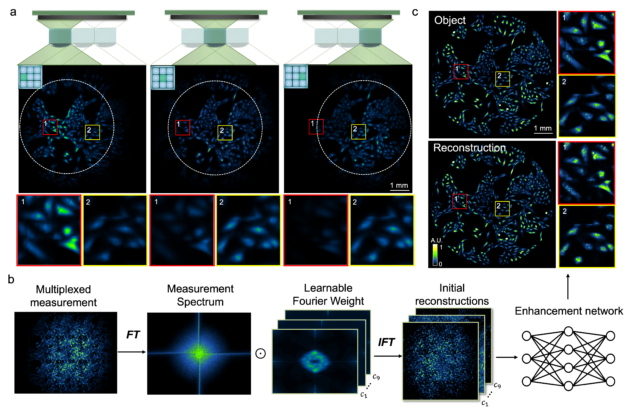
Wide-field, high-resolution reconstruction in computational multi-aperture miniscope using a Fourier neural network
Qianwan Yang, Ruipeng Guo, Guorong Hu, Yujia Xue, Yunzhe Li, and Lei Tian
Optica Vol. 11, Issue 6, pp. 860-871 (2024).
⭑ Github Project
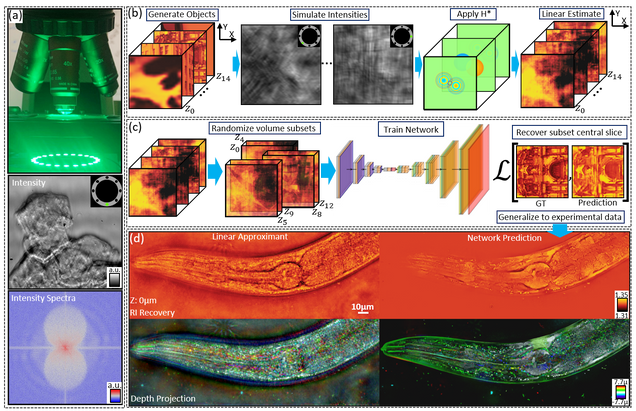
Multiple-scattering simulator-trained neural network for intensity diffraction tomography
A. Matlock, J. Zhu, L. Tian
Optics Express 31, 4094-4107 (2023)
Recovering 3D phase features of complex biological samples traditionally sacrifices computational efficiency and processing time for physical model accuracy and reconstruction quality. Here, we overcome this challenge using an approximant-guided deep learning framework in a high-speed intensity diffraction tomography system. Applying a physics model simulator-based learning strategy trained entirely on natural image datasets, we show our network can robustly reconstruct complex 3D biological samples. To achieve highly efficient training and prediction, we implement a lightweight 2D network structure that utilizes a multi-channel input for encoding the axial information. We demonstrate this framework on experimental measurements of weakly scattering epithelial buccal cells and strongly scattering C. elegans worms. We benchmark the network’s performance against a state-of-the-art multiple-scattering model-based iterative reconstruction algorithm. We highlight the network’s robustness by reconstructing dynamic samples from a living worm video. We further emphasize the network’s generalization capabilities by recovering algae samples imaged from different experimental setups. To assess the prediction quality, we develop a quantitative evaluation metric to show that our predictions are consistent with both multiple-scattering physics and experimental measurements.
Recovery of Continuous 3D Refractive Index Maps from Discrete Intensity-Only Measurements using Neural Fields
Renhao Liu, Yu Sun, Jiabei Zhu, Lei Tian, Ulugbek Kamilov
Nature Machine Intelligence 4, 781–791 (2022).
Intensity diffraction tomography (IDT) refers to a class of optical microscopy techniques for imaging the three-dimensional refractive index (RI) distribution of a sample from a set of two-dimensional intensity-only measurements. The reconstruction of artefact-free RI maps is a fundamental challenge in IDT due to the loss of phase information and the missing-cone problem. Neural fields has recently emerged as a new deep learning approach for learning continuous representations of physical fields. The technique uses a coordinate-based neural network to represent the field by mapping the spatial coordinates to the corresponding physical quantities, in our case the complex-valued refractive index values. We present Deep Continuous Artefact-free RI Field (DeCAF) as a neural-fields-based IDT method that can learn a high-quality continuous representation of a RI volume from its intensity-only and limited-angle measurements. The representation in DeCAF is learned directly from the measurements of the test sample by using the IDT forward model without any ground-truth RI maps. We qualitatively and quantitatively evaluate DeCAF on the simulated and experimental biological samples. Our results show that DeCAF can generate high-contrast and artefact-free RI maps and lead to an up to 2.1-fold reduction in the mean squared error over existing methods.
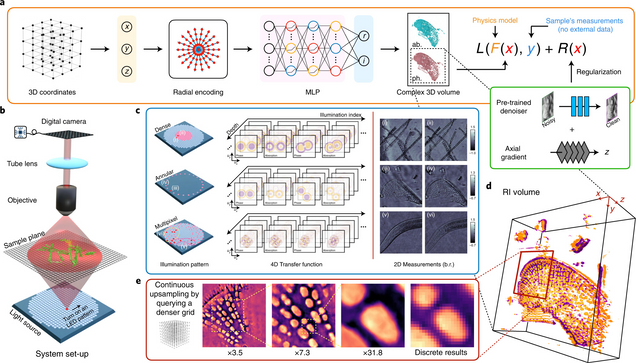
Deep learning-augmented Computational Miniature Mesoscope
Yujia Xue, Qianwan Yang, Guorong Hu, Kehan Guo, Lei Tian
Optica 9, 1009-1021 (2022)
Fluorescence microscopy is essential to study biological structures and dynamics. However, existing systems suffer from a tradeoff between field-of-view (FOV), resolution, and complexity, and thus cannot fulfill the emerging need of miniaturized platforms providing micron-scale resolution across centimeter-scale FOVs. To overcome this challenge, we developed Computational Miniature Mesoscope (CM
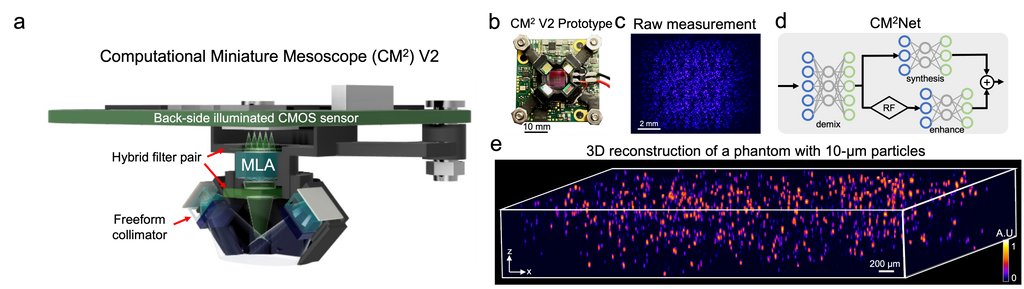
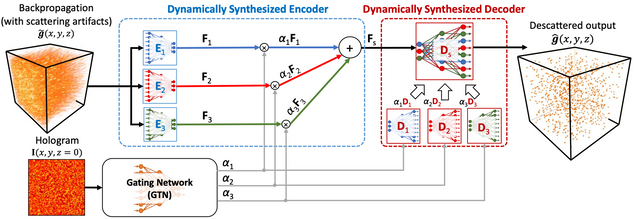
Adaptive 3D descattering with a dynamic synthesis network
Waleed Tahir, Hao Wang, Lei Tian
Light: Science & Applications 11, 42, 2022
Deep learning has been broadly applied to imaging in scattering applications. A common framework is to train a “descattering” neural network for image recovery by removing scattering artifacts. To achieve the best results on a broad spectrum of scattering conditions, individual “expert” networks have to be trained for each condition. However, the performance of the expert sharply degrades when the scattering level at the testing time differs from the training. An alternative approach is to train a “generalist” network using data from a variety of scattering conditions. However, the generalist generally suffers from worse performance as compared to the expert trained for each scattering condition. Here, we develop a drastically different approach, termed dynamic synthesis network (DSN), that can dynamically adjust the model weights and adapt to different scattering conditions. The adaptability is achieved by a novel architecture that enables dynamically synthesizing a network by blending multiple experts using a gating network. Notably, our DSN adaptively removes scattering artifacts across a continuum of scattering conditions regardless of whether the condition has been used for the training, and consistently outperforms the generalist. By training the DSN entirely on a multiple-scattering simulator, we experimentally demonstrate the network’s adaptability and robustness for 3D descattering in holographic 3D particle imaging. We expect the same concept can be adapted to many other imaging applications, such as denoising, and imaging through scattering media. Broadly, our dynamic synthesis framework opens up a new paradigm for designing highly adaptive deep learning and computational imaging techniques.
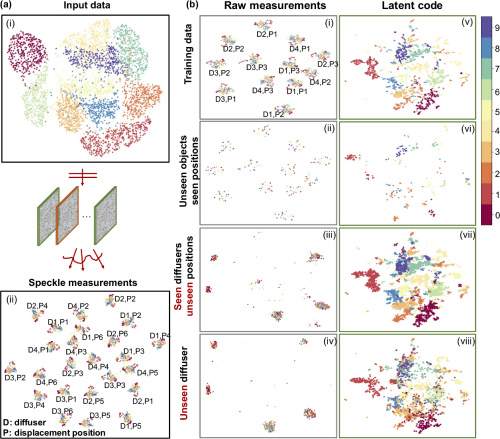
Displacement-agnostic coherent imaging through scatter with an interpretable deep neural network
Y. Li, S. Cheng, Y. Xue, L. Tian
Optics Express Vol. 29, Issue 2, pp. 2244-2257 (2021).
Coherent imaging through scatter is a challenging task. Both model-based and data-driven approaches have been explored to solve the inverse scattering problem. In our previous work, we have shown that a deep learning approach can make high-quality and highly generalizable predictions through unseen diffusers. Here, we propose a new deep neural network model that is agnostic to a broader class of perturbations including scatterer change, displacements, and system defocus up to 10× depth of field. In addition, we develop a new analysis framework for interpreting the mechanism of our deep learning model and visualizing its generalizability based on an unsupervised dimension reduction technique. We show that our model can unmix the scattering-specific information and extract the object-specific information and achieve generalization under different scattering conditions. Our work paves the way to a robust and interpretable deep learning approach to imaging through scattering media.
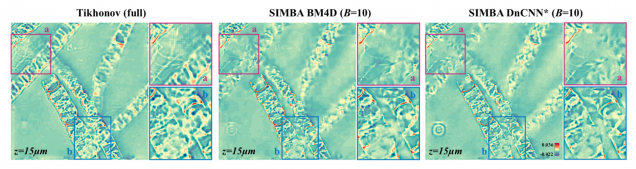
SIMBA: Scalable Inversion in Optical Tomography using Deep Denoising Priors
Zihui Wu, Yu Sun, Alex Matlock, Jiaming Liu, Lei Tian, Ulugbek S. Kamilov
IEEE Journal of Selected Topics in Signal Processing 14(6), 2020.

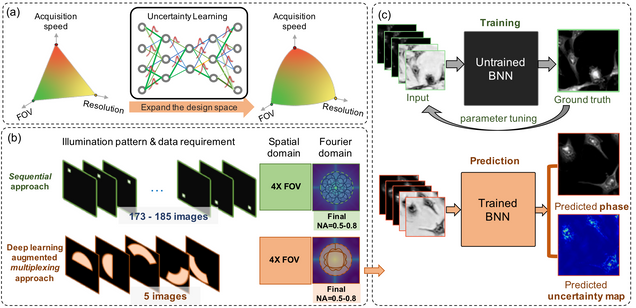
Reliable deep learning-based phase imaging with uncertainty quantification
Yujia Xue, Shiyi Cheng, Yunzhe Li, Lei Tian
Optica 6, 618-629 (2019).
Emerging deep-learning (DL)-based techniques have significant potential to revolutionize biomedical imaging. However, one outstanding challenge is the lack of reliability assessment in the DL predictions, whose errors are commonly revealed only in hindsight. Here, we propose a new Bayesian convolutional neural network (BNN)-based framework that overcomes this issue by quantifying the uncertainty of DL predictions. Foremost, we show that BNN-predicted uncertainty maps provide surrogate estimates of the true error from the network model and measurement itself. The uncertainty maps characterize imperfections often unknown in real-world applications, such as noise, model error, incomplete training data, and out-of-distribution testing data. Quantifying this uncertainty provides a per-pixel estimate of the confidence level of the DL prediction as well as the quality of the model and data set. We demonstrate this framework in the application of large space–bandwidth product phase imaging using a physics-guided coded illumination scheme. From only five multiplexed illumination measurements, our BNN predicts gigapixel phase images in both static and dynamic biological samples with quantitative credibility assessment. Furthermore, we show that low-certainty regions can identify spatially and temporally rare biological phenomena. We believe our uncertainty learning framework is widely applicable to many DL-based biomedical imaging techniques for assessing the reliability of DL predictions.
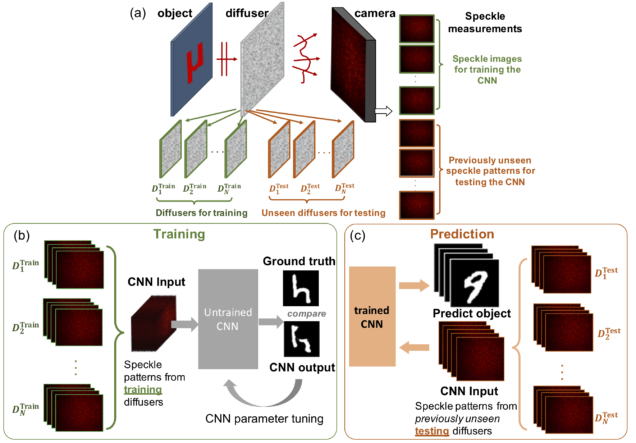
Deep speckle correlation: a deep learning approach towards scalable imaging through scattering media
Yunzhe Li, Yujia Xue, Lei Tian
Optica 5, 1181-1190 (2018).
⭑ Top 5 most cited articles in Optica published in 2018 (Source: Google Scholar)
Imaging through scattering is an important yet challenging problem. Tremendous progress has been made by exploiting the deterministic input–output “transmission matrix” for a fixed medium. However, this “one-to-one” mapping is highly susceptible to speckle decorrelations – small perturbations to the scattering medium lead to model errors and severe degradation of the imaging performance. Our goal here is to develop a new framework that is highly scalable to both medium perturbations and measurement requirement. To do so, we propose a statistical “one-to-all” deep learning (DL) technique that encapsulates a wide range of statistical variations for the model to be resilient to speckle decorrelations. Specifically, we develop a convolutional neural network (CNN) that is able to learn the statistical information contained in the speckle intensity patterns captured on a set of diffusers having the same macroscopic parameter. We then show for the first time, to the best of our knowledge, that the trained CNN is able to generalize and make high-quality object predictions through an entirely different set of diffusers of the same class. Our work paves the way to a highly scalable DL approach for imaging through scattering media.