Revealing hidden signatures
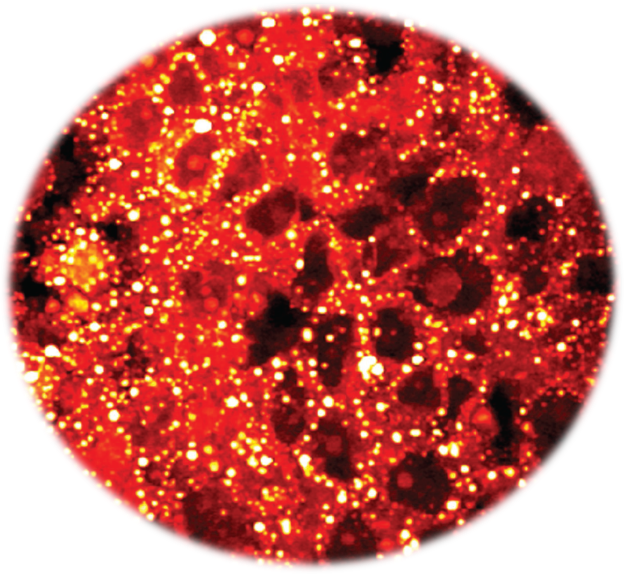
2.1 Cancer metabolic signatures
By label-free spectroscopic imaging of patient specimens, our team discovered an aberrant storage of cholesteryl ester in aggressive human prostate cancer (Cell Metabolism 2014). This study identified a novel target that can be used to treat late-stage human prostate cancer (US patent 9,164,084 B2) through a nanomedicine formulation (ACS Nano 2015). More recently, we identified lipid unsaturation as a metabolic marker of ovarian cancer stem cells (Cell Stem Cell 2017), which paves the foundation for suppression of cancer stem cells.
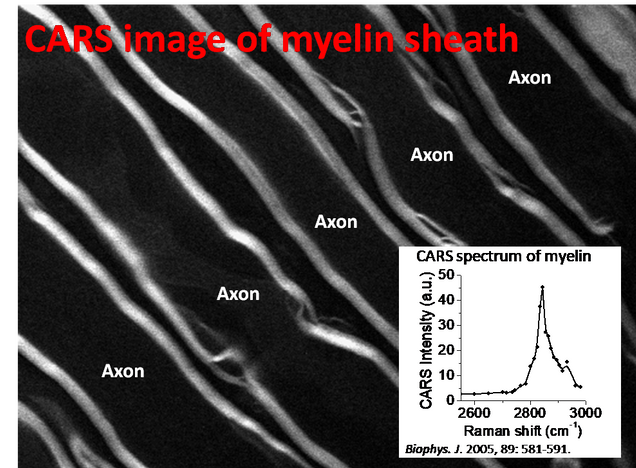
2.2 Myelin sheath de-/re-generation
The Cheng laboratory was the first to show label-free CARS imaging of myelin sheath in its natural state [Biophys J 2005, 89: 581-591], now pursued by different groups. This work led to Cheng group’s first R01 grant (2005-2008), and triggered the development of a nanomedicine for functional recovery of an injured spinal cord [Nature Nanotechnology 2010, 5: 80-87], supported by a translational research award from the Coulter Foundation.
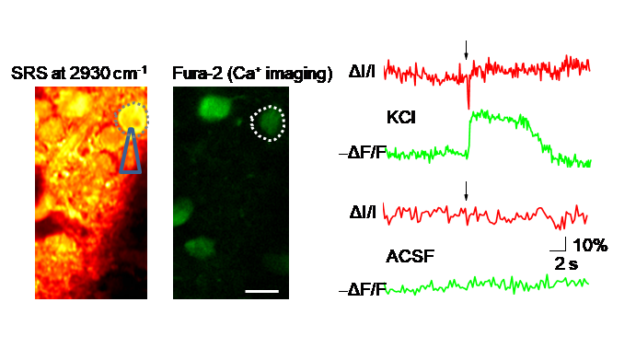
2.3 Spectroscopic imaging of membrane potential in living neurons
Detecting membrane potentials is critical for understanding how neuronal networks process information. We reported a vibrational spectroscopic signature of neuronal membrane potentials identified through hyperspectral stimulated Raman scattering (SRS) imaging of patched primary neurons [JPC Letters 2017]. High-speed SRS imaging allowed direct visualization of puff-induced depolarization of multiple neurons in mouse brain slices, confirmed by simultaneous calcium imaging. By implementing a dual-SRS balanced detection scheme, we detected single action potentials in electrically stimulated neurons. By pre-resonance SRS, we recently demonstrated spectroscopic detection of voltage-sensitive rhodopsins [JPC Letters 2019].
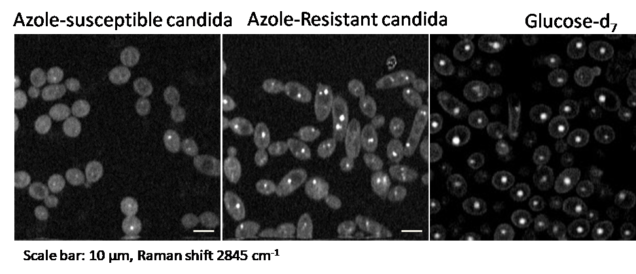
2.4 Antibiotic susceptibility determination within one cell cycle at single bacterium level
The widespread use of antibiotics has significantly increased the number of resistant bacteria, which has also increased the urgency of rapid bacterial detection and profiling their antibiotics response. Current methods for antibiotic susceptibility testing (AST) require at least 16 to 24 h to conduct. Therefore, there is an urgent need for a rapid method that can test the susceptibility of bacteria in a culture-free manner. We demonstrated a rapid AST method by monitoring the glucose metabolic activity of live bacteria at the single cell level with hyperspectral stimulated Raman scattering imaging of and fungal cells [Anal Chem 2017] and bacteria [Anal Chem 2018].