Research
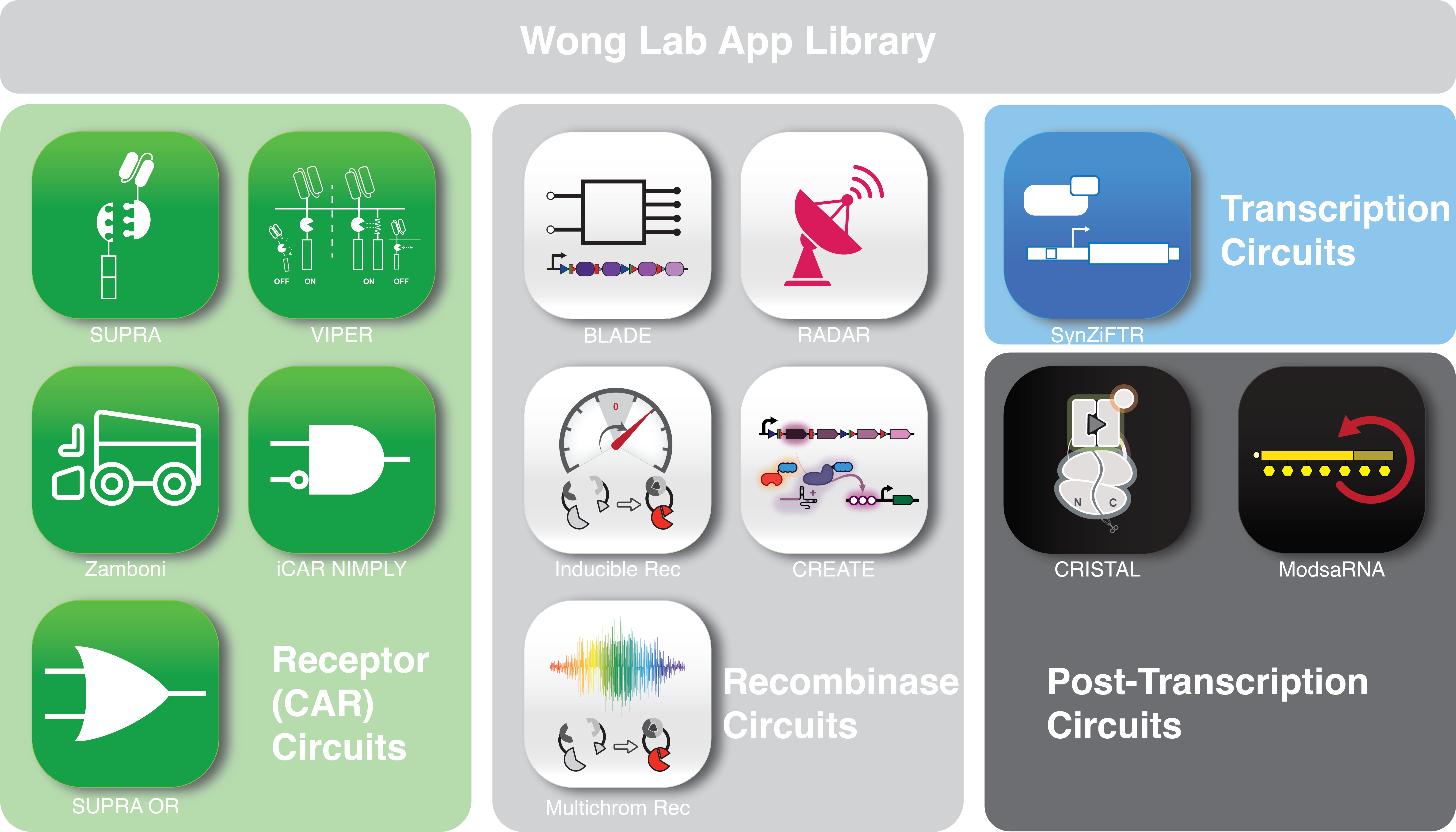
The Wong lab is a DESIGN-driven lab where we apply synthetic biology to engineer desired tools (Apps) and properties in mammalian cells, with application in understanding cellular design principles and developing cellular therapy. Our research program can be broadly categorized into several areas: (1) Advanced chimeric antigen receptor (CAR) design, (2) Drug-inducible gene and signaling switches, (3) Biocomputer engineering, (4) self-amplifying RNA. Please see our publications for more information.